In this lesson we’ll explore AWS SageMaker data processing jobs and how they help offload heavy data preparation from your interactive notebook environment. Processing jobs allow you to run preprocessing at scale in dedicated compute, improving speed, reproducibility, and maintainability. What we’ll cover:Documentation Index
Fetch the complete documentation index at: https://notes.kodekloud.com/llms.txt
Use this file to discover all available pages before exploring further.
- Why running heavy preprocessing inside Jupyter notebooks is a problem
- How SageMaker processing jobs offload work to appropriately sized compute
- An end-to-end SKLearn example using the SageMaker Python SDK
- Processor classes and when to use each
- How to monitor processing jobs and their benefits
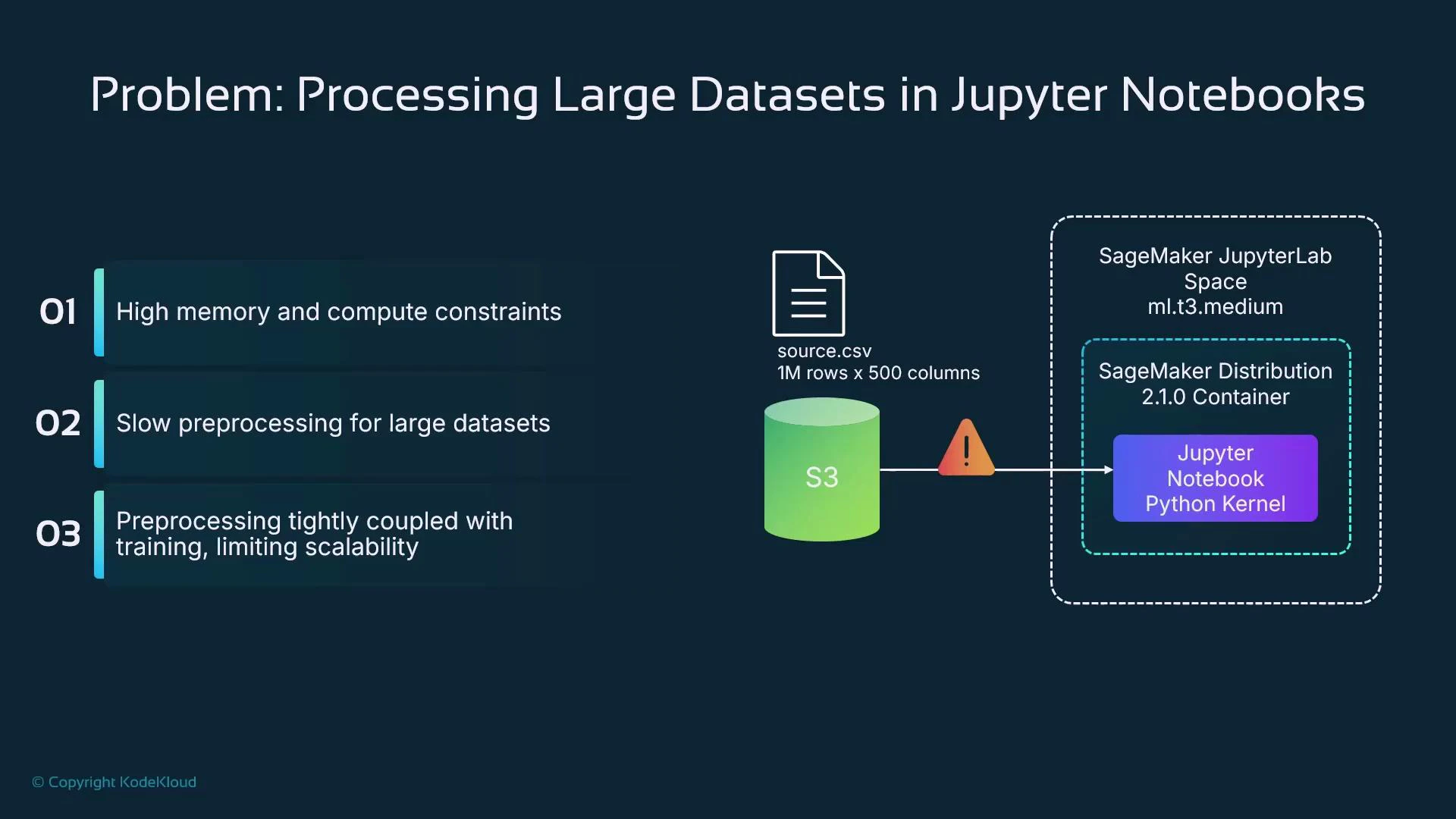
Problem context
Running data preparation inside a managed notebook (SageMaker Studio or Notebook Instances) ties the work to a fixed, often underpowered, instance type (for example, ml.t3.medium). Large datasets — for example, a CSV with 1,000,000 rows and 500 columns — frequently require far more CPU, memory, or distributed processing than a notebook kernel can provide. Common heavy preprocessing tasks include:- Missing-value imputation
- Numeric scaling (StandardScaler, MinMaxScaler)
- One-hot encoding categorical variables
- Feature engineering (arithmetic combinations, time differences)
- Train/validation splits with class balancing
- Sampling, SMOTE, or other resampling approaches
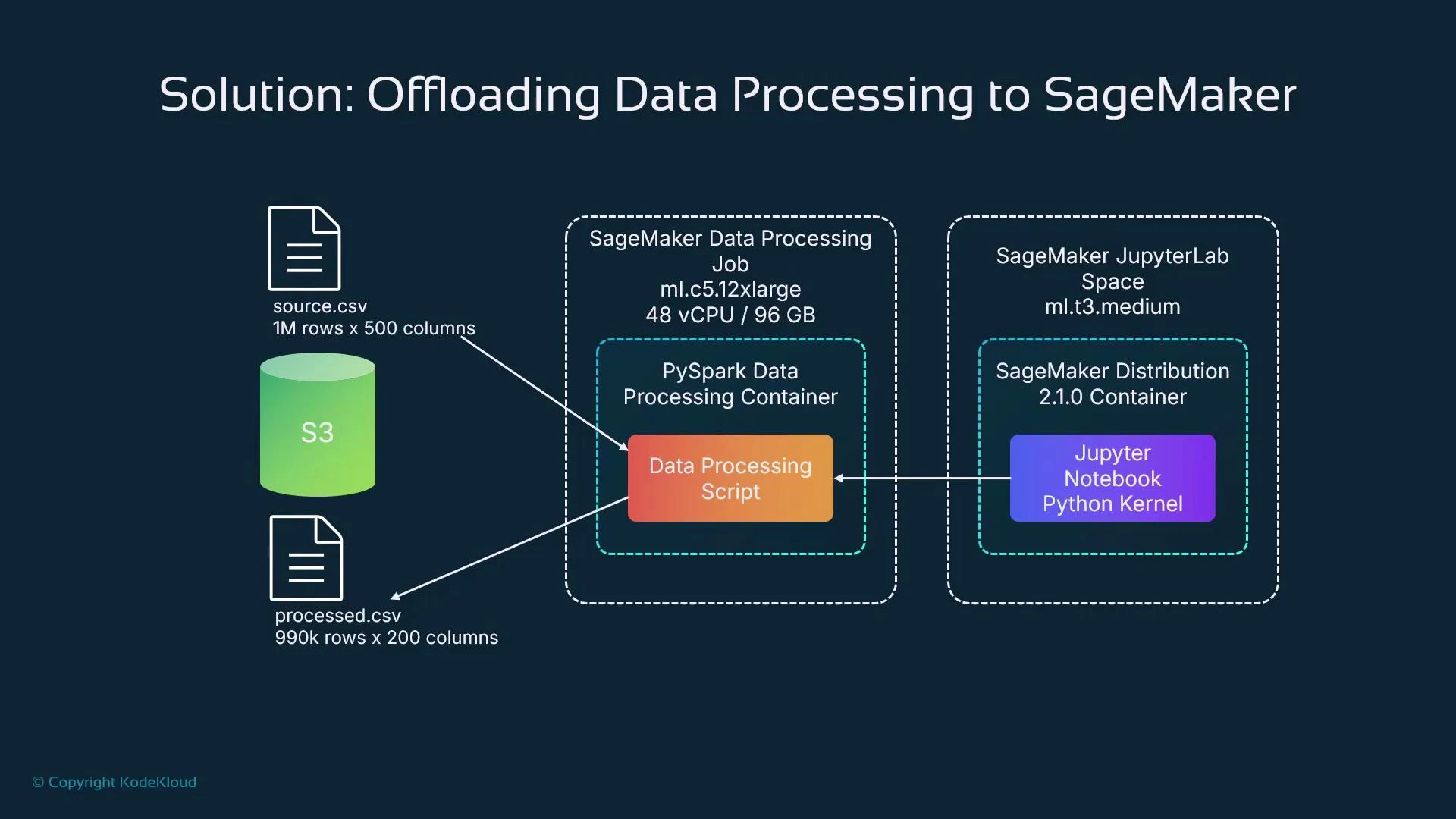
Solution: Offload preprocessing to SageMaker processing jobs
SageMaker processing jobs run the provided script inside managed containers on dedicated instances or clusters you request. From a notebook, you submit a processing job (via the SageMaker Python SDK or APIs) and SageMaker provisions the requested compute, pulls the container, executes the script, and writes outputs to S3. Typical options:- Single large instance (e.g., ml.c5.12xlarge) for CPU-bound tasks
- Multi-node clusters with PySpark for distributed workloads
- GPU instances for GPU-accelerated preprocessing (image feature extraction, large model-based transforms)
- Managed framework containers (SKLearn, PySpark, PyTorch, TensorFlow) or a custom container via ScriptProcessor
- Notebook defines processing job: code, inputs, outputs, compute resources.
- Notebook submits the job using the SageMaker SDK.
- SageMaker provisions instance(s), pulls the container image, runs the code, and writes outputs to S3.
- Notebook (or pipeline) consumes processed artifacts (S3, Feature Store, or downstream training).
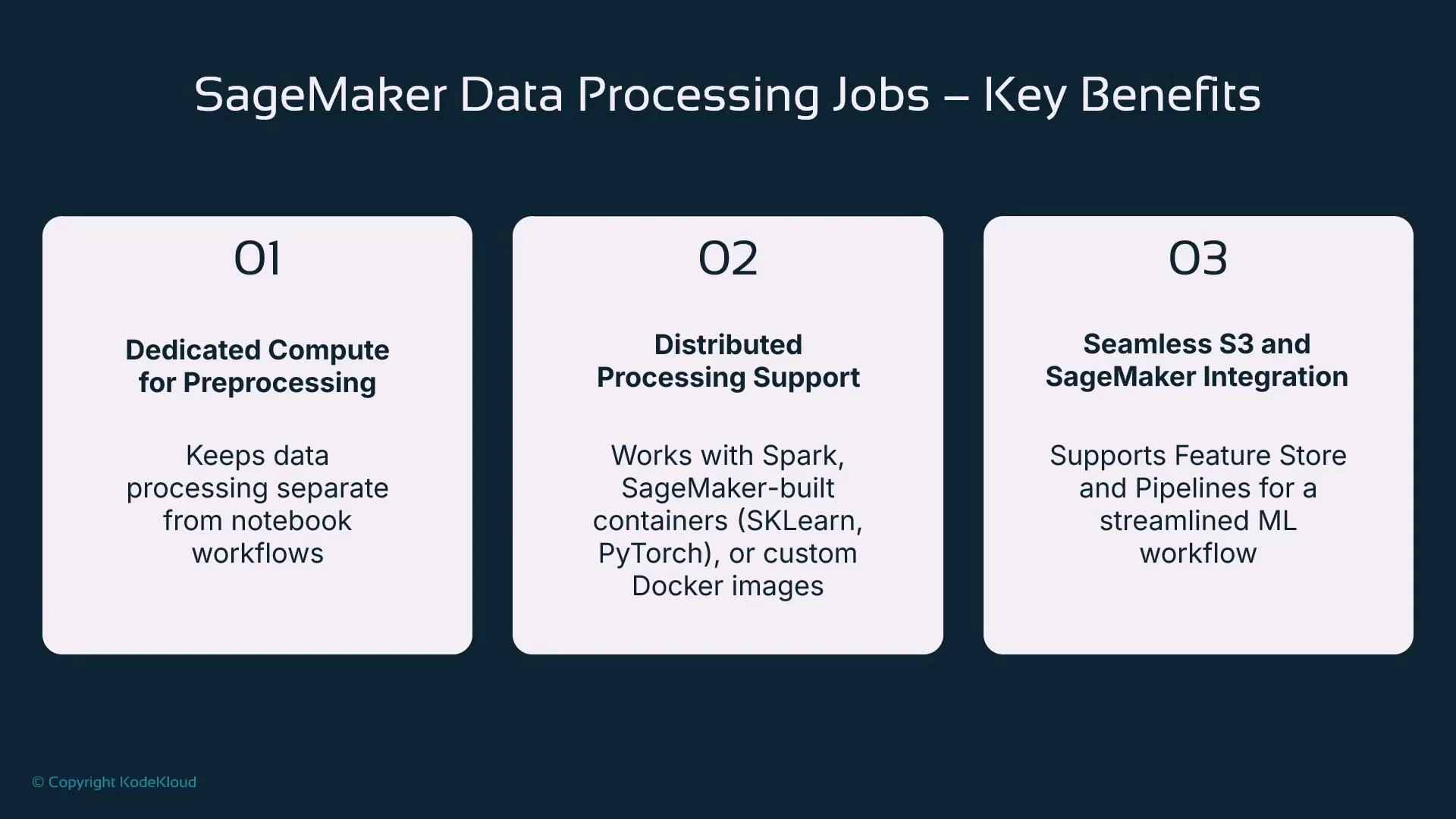
- Decouple preprocessing compute from interactive development sessions
- Right-size compute for each job (single large instance vs. multi-node clusters)
- Reuse managed framework containers when available (scikit-learn, PySpark, PyTorch, TensorFlow)
- Bring your own container for custom dependencies and runtime control (ScriptProcessor)
- Integrate with S3, SageMaker Feature Store, and SageMaker Pipelines for reproducible workflows
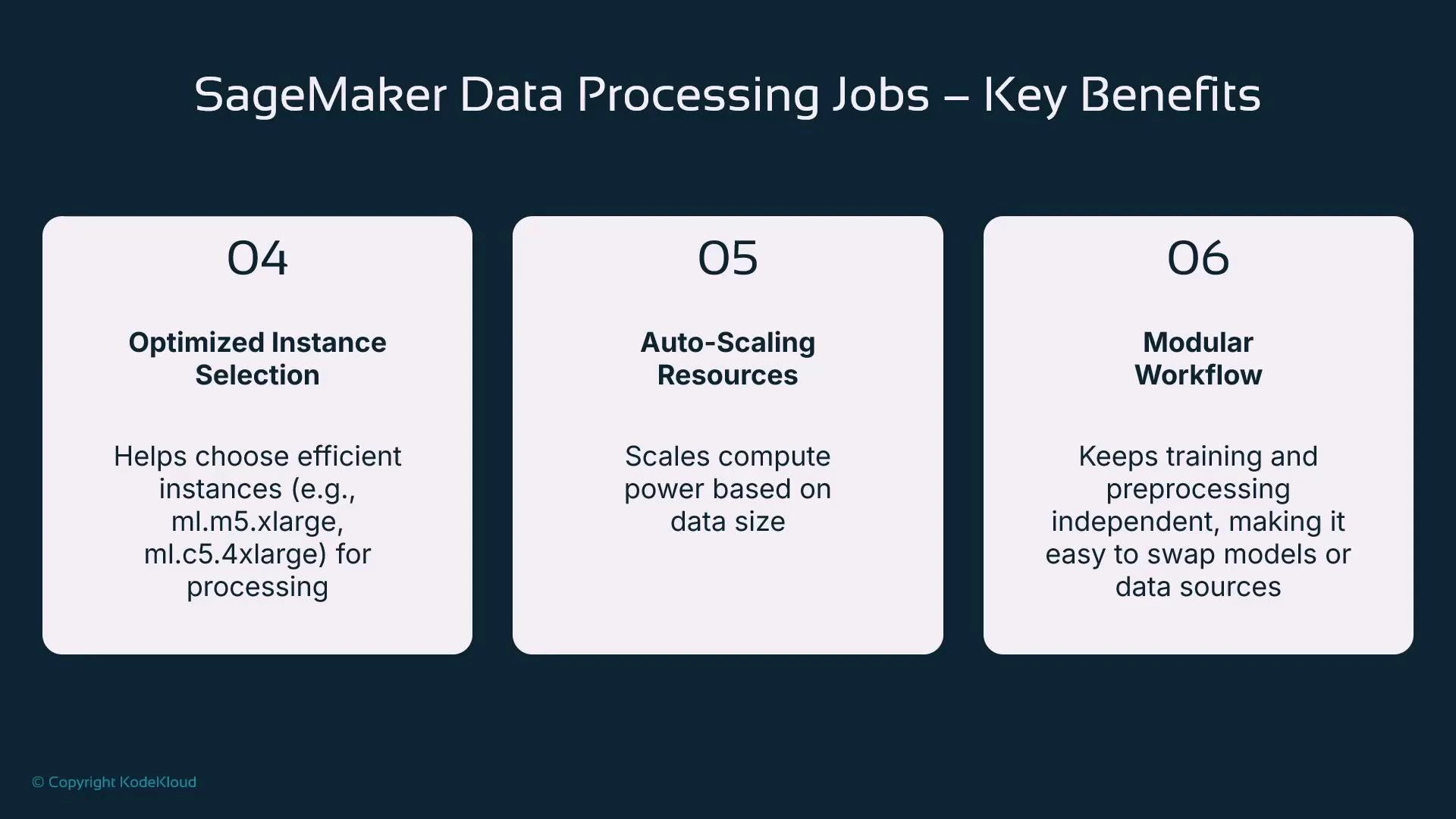
Right-sizing and scaling guidance
- Choose instance type and count based on CPU, memory, and I/O needs (single large vs. multi-node).
- Use PySparkProcessor for scale-out and parallelism across nodes.
- Prefer a powerful instance for a short duration rather than a weak instance for a long run to reduce elapsed time and often cost.
- Modularize preprocessing into discrete jobs so they can be versioned, retried, and reused independently by teams.
Example: SKLearn processing job (inline script)
This concise SKLearn example demonstrates writing a small preprocessing script, configuring an SKLearnProcessor, and submitting a processing job. Adjust S3 URIs, roles, instance types, and framework versions for your environment.- The script runs inside the managed container on the provisioned instance.
- Processing containers expect input/output under /opt/ml/processing/input and /opt/ml/processing/output.
- SKLearnProcessor chooses a managed scikit-learn image that matches the framework_version.
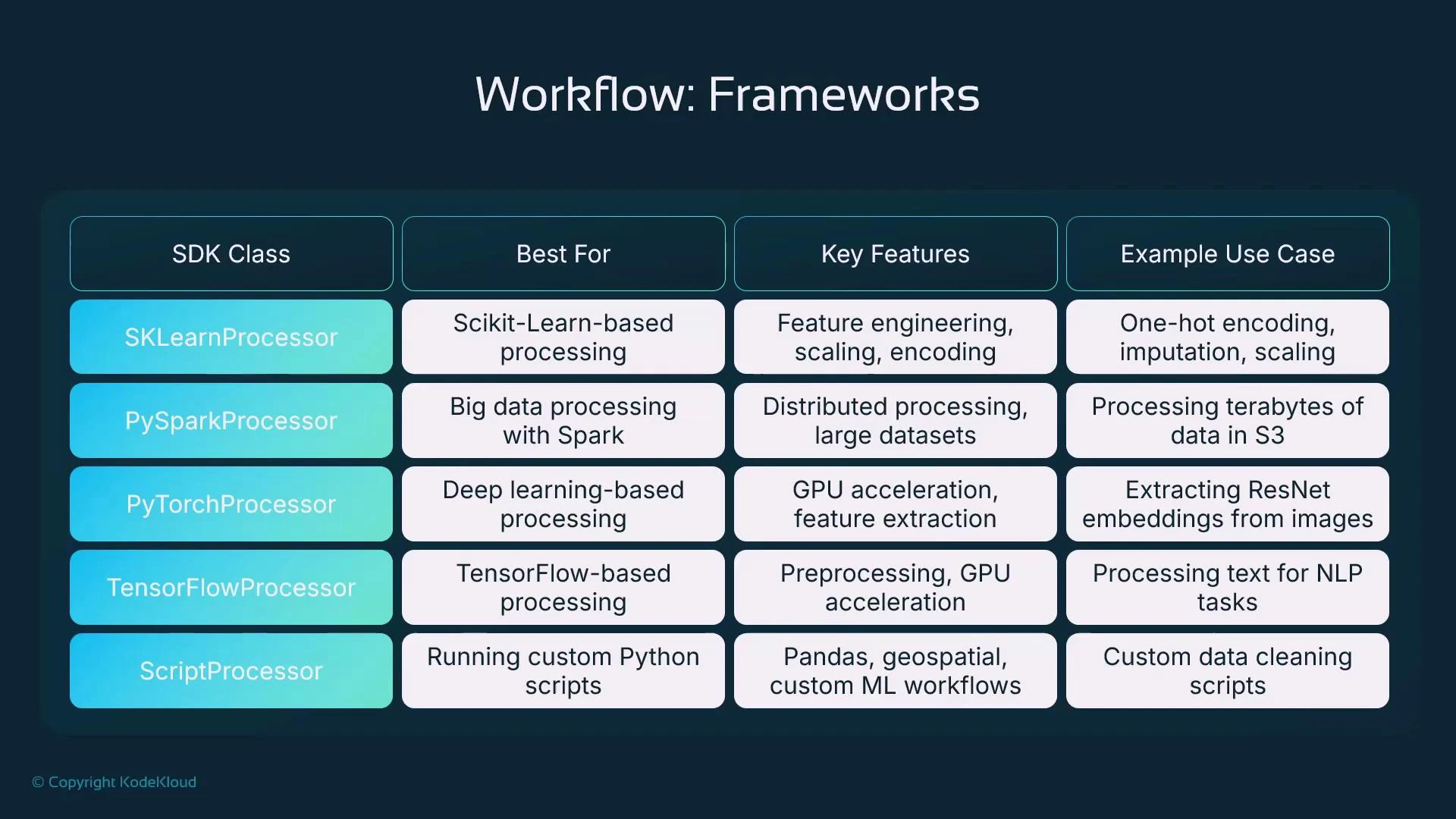
Processor classes — when to use each
The SageMaker SDK exposes processor classes that map to common use cases. This table helps choose the right processor:| Processor class | Use case | Example |
|---|---|---|
| SKLearnProcessor | Tabular preprocessing, feature engineering, scaling, imputation | Use for pandas + scikit-learn transforms |
| PySparkProcessor | Large-scale distributed ETL using Spark | Multi-node transformations, aggregations, joins |
| PyTorchProcessor | GPU-accelerated preprocessing for images or model-based transforms | Image feature extraction with pretrained models |
| TensorFlowProcessor | TensorFlow-based preprocessing or GPU tasks | Text sequence transforms or TF-based feature ops |
| ScriptProcessor | Custom dependencies or runtime — provide an ECR image | Bring-your-own Docker image for specialized libs |
ScriptProcessor (custom container) example
Use ScriptProcessor when you require a custom image in ECR.When using ScriptProcessor with a custom ECR image, ensure the execution role has permission to pull from ECR and that the image is compatible with SageMaker’s processing lifecycle. Also review instance costs for large or GPU-powered instances.
PyTorch processor (GPU) example
Use for GPU-accelerated preprocessing like image feature extraction:PySpark processor (multi-node) example
Use for distributed ETL and scale-out across nodes:Monitoring processing jobs
You can monitor jobs in the AWS Console (SageMaker > Processing) to view job status, container image, role, entry point script, timestamps, and logs. Logs are available via CloudWatch — streaming them during runs is helpful for debugging.You can stream processing job logs via CloudWatch Logs. Open the Processing job details in the console to find the CloudWatch log group and observe real-time output for troubleshooting.
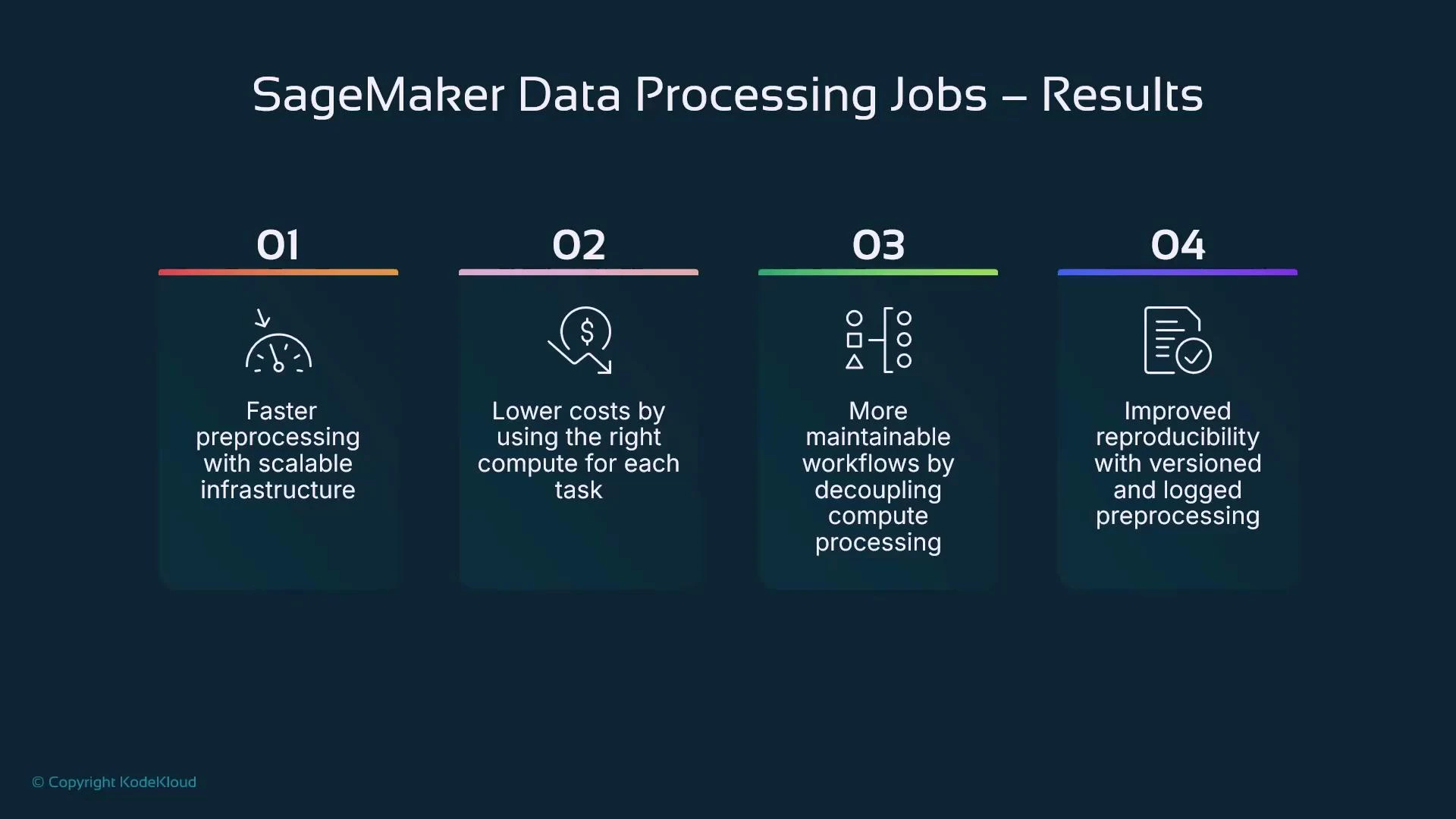
Benefits recap
- Faster preprocessing using scalable, dedicated compute
- Lower operational cost by right-sizing compute for each task
- More maintainable workflows by decoupling preprocessing from interactive sessions
- Improved reproducibility via versioned scripts, container images, and logged runs
- Use processing jobs to delegate heavy preprocessing to managed containers and appropriately sized compute.
- Choose SKLearn/PySpark/PyTorch/TensorFlow processors for common frameworks; choose ScriptProcessor when you need a custom container.
- Scale out using PySparkProcessor (instance_count > 1) for distributed workloads.
- Inside containers use /opt/ml/processing/input and /opt/ml/processing/output; inputs and outputs are typically S3 URIs.
- Monitor jobs in SageMaker console and CloudWatch for auditing and debugging.
- Modular, version-controlled preprocessing improves reproducibility across teams.
Links and references
- SageMaker processing jobs documentation
- SageMaker Python SDK documentation
- Amazon S3
- AWS CloudWatch Logs