This lesson explains what Jupyter Notebooks solve, why data scientists favor them for experimentation and collaboration, how to run them (hosted services or locally), and their primary benefits. The dominant language for data science is Python. To develop models and analyze data you need an appropriate Python environment. Several options exist, each with trade-offs depending on whether your goal is quick experimentation, reproducible analysis, or production-quality development.Documentation Index
Fetch the complete documentation index at: https://notes.kodekloud.com/llms.txt
Use this file to discover all available pages before exploring further.
Problem: Python REPL (interactive shell)
A minimal starting point is the Python interactive shell (a REPL — Read, Evaluate, Print Loop). If Python is installed locally, runningpython opens a prompt where you can try commands like print("hello") and get immediate results. This is great for quick experiments and learning, but it doesn’t scale for multiline, reusable, or version-controlled code. For moderate-length scripts (tens or hundreds of lines) you typically use Python script files and an editor or IDE instead.

Alternatives: IDEs
Integrated development environments (IDEs) supply many productivity features that help when building applications:- Code completion and parameter hints
- Syntax highlighting and linting
- Debugging (breakpoints, step over/into, variable watches)
- AI-assisted suggestions in some editors

Why data scientists need something different
Data exploration typically requires multiple tools and rich visualizations to understand feature distributions and correlations. Libraries such as Matplotlib and Seaborn are commonly used to create charts that drive feature engineering and model design. REPLs and traditional IDEs can be limiting for reproducibility and collaborative experiments. To let colleagues reproduce your analysis or inspect “how you got that result,” you need an environment that preserves code, outputs (including charts), and narrative explanation together.
Solution: Jupyter Notebooks
Jupyter Notebooks are an interactive, web-based environment designed for experimentation and sharing. Key features:- Code organized into cells so you can run small chunks independently.
- Inline outputs: text, tables, and visualizations display directly below code cells.
- Persistence of inputs and outputs for reproducible experiments.
- Markdown cells let you add narrative, headings, and LaTeX math for clear documentation.

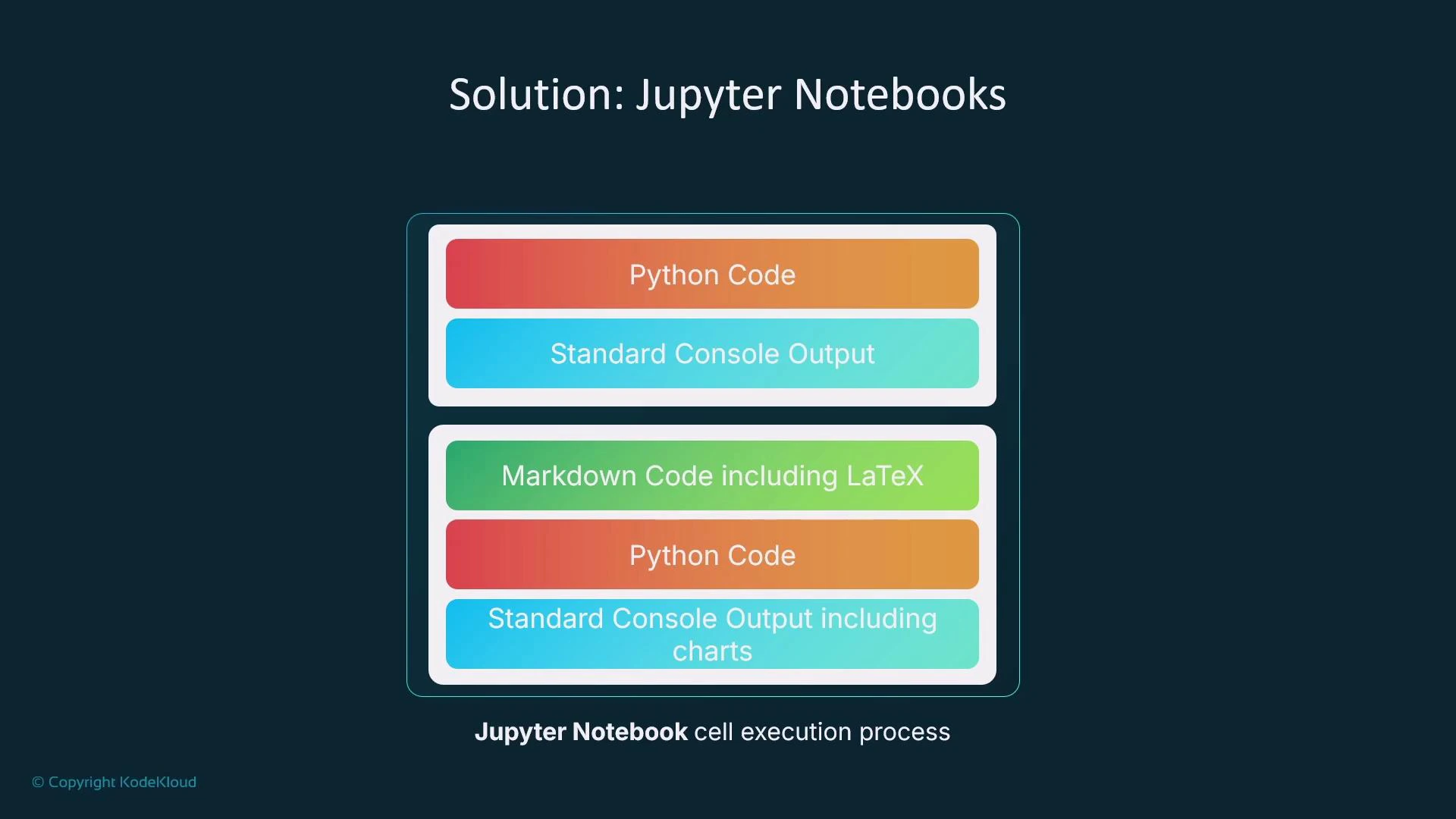
Notebook structure and cells
Notebooks contain two primary cell types:- Code cells: execute Python and display standard output inline. Visualizations from Matplotlib or Seaborn are rendered directly and saved with the notebook.
- Markdown cells: document steps, reasoning, and include LaTeX-compatible math.

Example — interactive notebook cells
A short example illustrating two code cells and their outputs. When executed in a notebook the printed results appear inline and are persisted with the .ipynb file.Comparison with other Python environments
Jupyter Notebooks combine interactive execution, inline visualization, and narrative markdown — making them ideal for data exploration, reproducibility, and teaching. REPLs are interactive but lack integrated visualization and reproducibility features; IDEs excel at debugging, code navigation, and production development, but are less naturally suited to exploratory, step-by-step scientific workflows.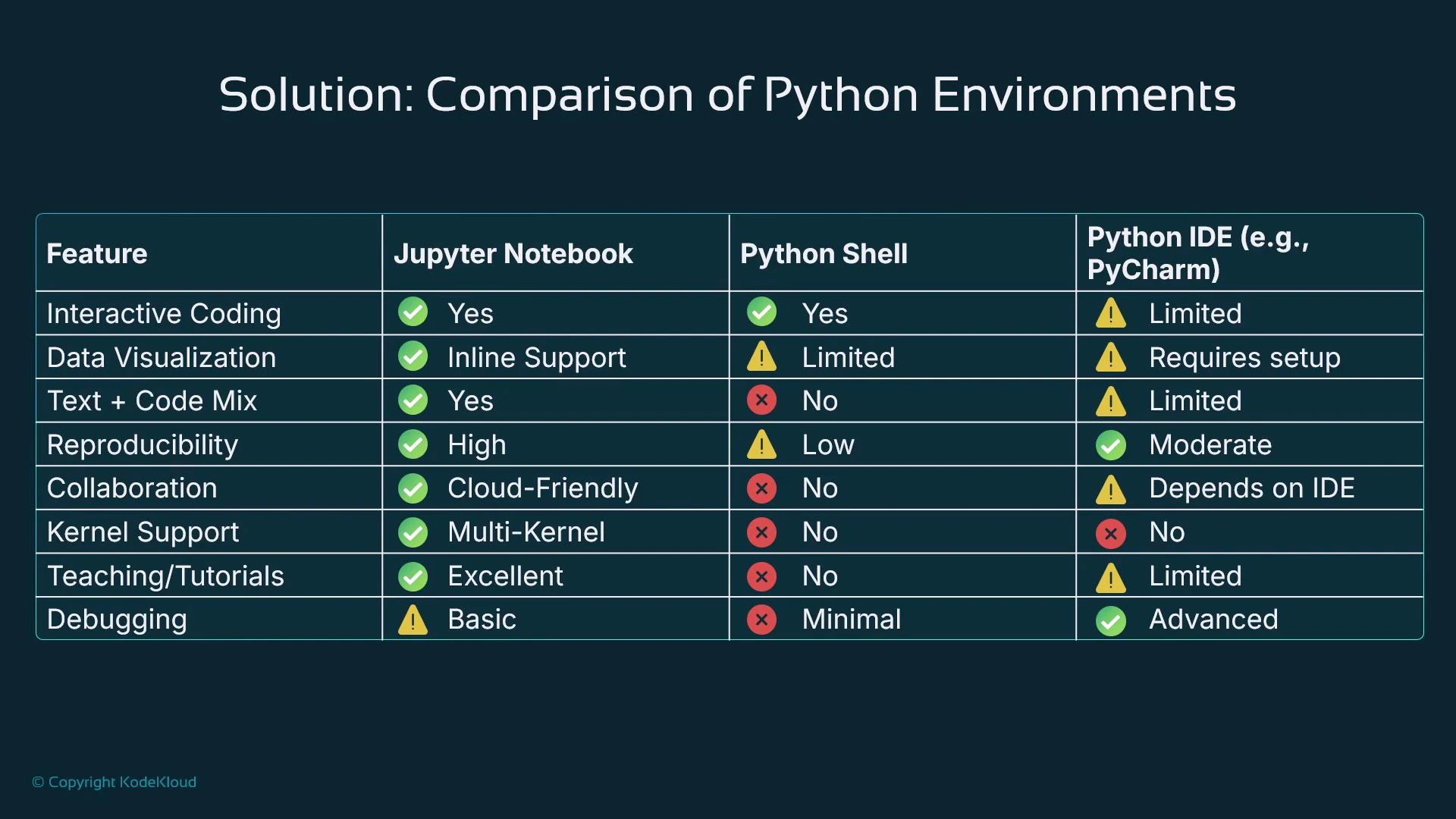
| Resource Type | Best for | Strengths |
|---|---|---|
| Jupyter Notebook / JupyterLab | Exploratory data analysis, visualization, reproducible research, teaching | Inline plots, markdown + code, shareable .ipynb, easy iteration |
| Python REPL (interactive shell) | Quick experiments, learning | Immediate feedback, minimal overhead |
| Python IDE (VS Code, PyCharm) | Production apps, debugging, refactoring | Advanced debugging, linting, project tooling, scalability |
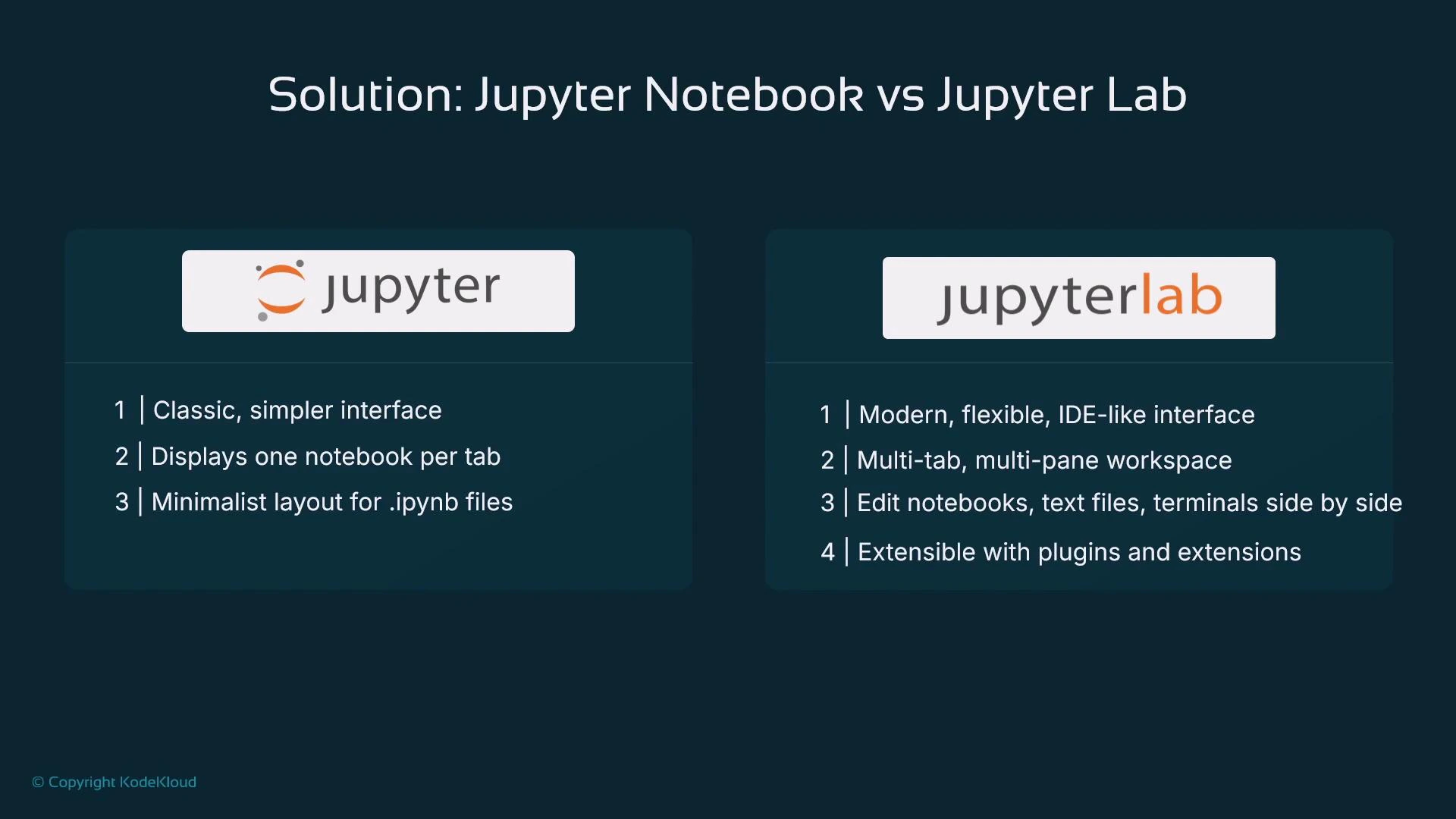
Jupyter vs JupyterLab
Two common interfaces:- Jupyter (classic): a simpler UI focused on one notebook at a time. Files use the .ipynb extension.
- JupyterLab: a modern, extensible IDE-like multi-pane interface. Supports multiple open files, terminals, and many extensions (Git integration, linters, documentation tools, vendor plugins).

Example: install packages from a notebook-attached terminal
Files, version control, and collaboration
Notebooks are saved as .ipynb files that embed code cells, markdown cells, and the outputs produced by executed cells. For collaborative projects use version control (e.g., Git), but be mindful of large outputs and binary-encoded images included in notebooks.Best practice: Use markdown cells to explain intent and decisions, keep notebooks modular (one experiment per notebook or clear sections), and clear heavy outputs before committing. Consider tools like nbstripout or nbdime to produce cleaner diffs and reduce repository noise.
Warning: Never commit notebooks that contain secrets or credentials. Also avoid large binary outputs (heavy plots, full datasets); store data externally and load it at runtime to keep repositories lightweight.
Summary
- Jupyter Notebooks provide an interactive, web-based environment ideal for exploratory data analysis, visualization, and reproducible experiments.
- They combine executable code, inline outputs (including charts), and rich markdown documentation to tell the story of your analysis.
- Choose JupyterLab for a multi-pane, extensible, IDE-like experience with integrated terminal and plugin support.
- Use traditional IDEs when you need advanced debugging, project organization, and production-ready development workflows.